Experiment reaches biology milestone with hard X-ray laser
Arizona State University builds protein beam injector; contributes nanocrystals
Unraveling the molecular basis of life is a long-standing quest of humanity. A breakthrough towards this goal was reported in a pair of studies published Feb. 3 in the scientific journal Nature, detailing a new method developed to determine structures of biomolecules based on diffraction from protein nanocrystals that are so small that they are not even visible under the microscope. A tiny aerojet nozzle provides a fully hydrated constant stream of nanocrystals, both supplied from an interdisciplinary research team at Arizona State University.
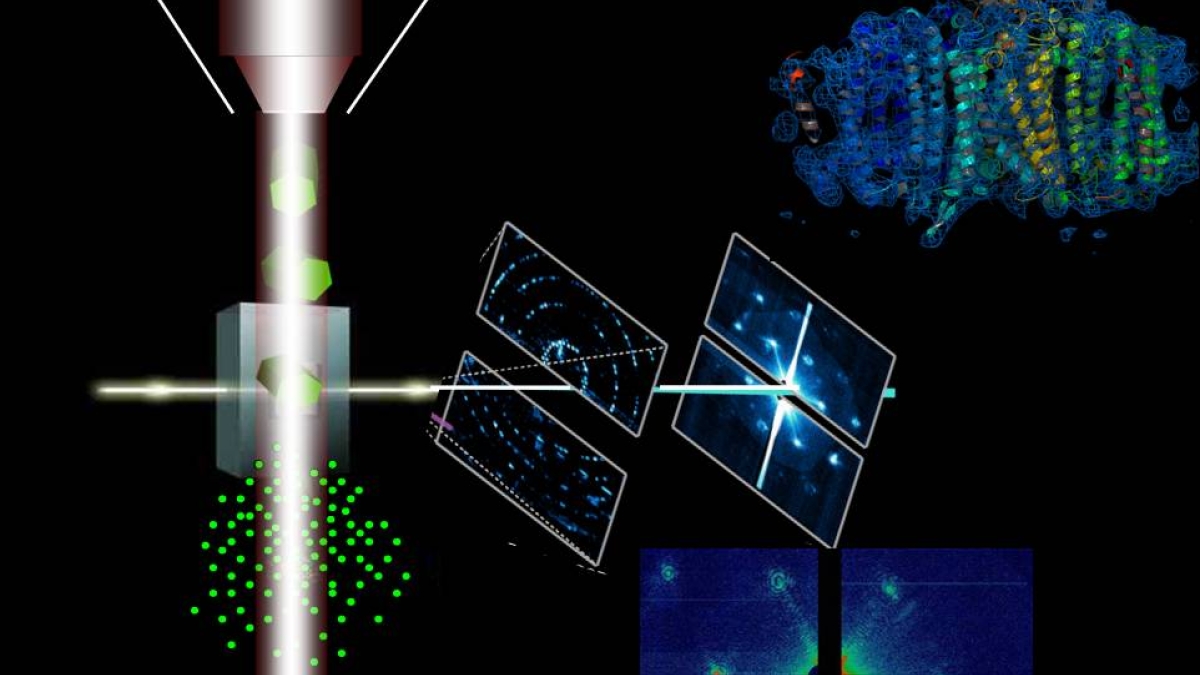
These developments allowed an international team of nearly 90 researchers to collect more than 3 million exquisitely clear diffraction pattern “snapshots” just before the crystals exploded in the extreme strong ultra-short femtosecond X-ray pulses that are so intense it will ultimately vaporize any solid material in its focus. The experiments show the great potential of the X-ray free-electron laser – the Linac Coherent Light Source located at the Department of Energy’s SLAC National Accelerator Laboratory – for new discoveries in biology and medicine.
“From the beginning, the resolution of images recorded by biologists has been limited by damage due to the radiation used,” said physicist John C.H. Spence, a Regents’ Professor in ASU’s College of Liberal Arts and Sciences, one of the lead authors of the studies. “But what happens if a pulse of imaging radiation is used that terminates before damage begins, yet contains sufficient photons to generate a useful scattering pattern?”
Many in the scientific community didn’t believe such a method could work. Yet, said Spence, “theory and recent experiments using soft X-rays had indicated that this might indeed provide a useful path to truly damage-free imaging, and then there was the recent invention of the hard X-ray laser (Linac Coherent Light Source).”
Unraveling protein structures
Living things are made up of biomolecules and proteins are the major constituents and "work horses" of all cells. “The unraveling of the structure and function of proteins is one of the most challenging goals in the post-genomic era,” said Petra Fromme, professor of chemistry and biochemistry at ASU, the second author of the nanocrystallography work.
Thirty percent of all proteins are not soluble but are embedded in membranes, she explained. “These membrane proteins, major players in the function of all living cells, are the targets for more than 50 percent of all drugs, yet, scientists have a lack of information on their structure and function. While more than 60,000 structures of soluble proteins have been determined to date only 256 membrane proteins structures are known and only four of them are human membrane proteins,” said Fromme, who directs ASU’s new Center for Membrane Proteins in Infectious Diseases, one of nine centers set up by the National Institutes of Health to decipher the structures of membrane proteins.
“This milestone research reported in Nature opens a new avenue for solving protein structures and will have a huge impact in a lot of areas, including development of clean energy and the medical field,” she said. “The determination of the protein structure will lead to the development of new drugs against cancer and infectious diseases, by manufacturing drugs that fit into the catalytic center of the proteins like a perfect key in a lock.”
For the nanocrystallography experiments, which took place in December 2009 at SLAC in California, Fromme’s lab at ASU supplied nanocrystals of photosystem I, a large membrane protein complex consisting of more than 100,000 atoms that acts as a biosolar energy converter in the process of oxygenic photosynthesis.
Protein crystals contain a lot of liquid and are “extremely weak, like a piece of butter in the sun,” Fromme explained. “When you just touch them, they’re destroyed.” It had taken her group 13 years to develop methods to grow the large, high-quality crystals of the photosystem I complex that allowed them to determine its molecular structure in 2001.
For the nanocrystallography experiment at the Linac Coherent Light Source, the research team used thousands of photosystem I nanocrystals instead. The significance of the experiment has the potential to revolutionize the field of protein crystallography and is reported in the Nature paper “Femtosecond X-ray protein nanocrystallography.”
"These experiments have the potential to be one of the most stunning scientific developments of the year" said Robert Nemanich, chair of ASU’s Department of Physics.
Little nozzle reaches target
Researcher Spence, with extensive experience in the physics of diffraction, led the ASU group that developed the protein beam injector, a tiny piece of equipment that fired a stream of protein solution to the pulsed hard X-ray beam at SLAC.
“The experimental run at the LCLS had exceeded all expectations,” said R. Bruce Doak, an ASU physics professor who has more than 30 years experience in the science of supersonic atomic and molecular beams. During the December 2009 experiment, he wrote: “The ASU liquid jet is performing flawlessly. We put together a dual reservoir supply system that allows us to switch samples without ever turning off the flow, and so the nozzle has now been running continuously for over 50 hours.
“Early on, we established the sample concentration of microcystallite photosystem I that delivers an X-ray diffraction pattern with almost every X-ray pulse – 30 diffraction images per second! No one here had ever seen anything like a 100% hit rat before, much less sustained for days; over 3 million stored diffraction images so far. The diffraction patterns are gorgeous,” wrote Doak, also an author on the Nature nanocrystallography paper.
Students are part of international effort
Other authors from ASU’s Department of Physics include Uwe Weierstall, a research professor; Kevin E. Schmidt, professor; Richard A. Kirian, doctoral student; and Xiaoyu Wang, doctoral student. Members of the team from ASU’s Department of Chemistry and Biochemistry include Mark S. Hunter, doctoral student; and faculty research associates Ingo Grotjohann and Raimund Fromme.
“Richard Kirian and Mark Hunter have worked for several years on the femtosecond crystallography project and are exemplars of outstanding graduate students at ASU, working in an interdisciplinary field,” said Petra Fromme. “Mark has grown all the photosystem I crystals for the experiments and developed new methods and technology for growth and biophysical characterization of protein nanocrystals in his PhD studies and Richard has developed new methods for evaluation and analysis of nanocrystallography data in his PhD studies.”
Heading up the international research effort and lead author on the nanocrystallography paper was Henry N. Chapman, DESY, Germany. Among others, the research team from more than 20 institutions included Janos Hajdu, M. Marvin Seibert, Tomas Ekeberg and Filipe R.N.C. Maia, Uppsala University, Sweden. Results of the second experiment performed the same week at SLAC are reported in the Nature paper “Single mimivirus particles intercepted and imaged with an X-ray laser.”
“These are early days for femtosecond diffractive imaging,” noted Spence, who provided the theory and much of the data analysis. “But first indications are that high-resolution data can now be obtained at the nanoscale by this method. If we can indeed ‘outrun’ the many radiation-damage processes in this way, it will open the way to future experiments on laser-excited samples, 3-D image reconstruction and a host of other experiments on fast imaging, all directed to the grand challenge of obtaining movies showing molecules at work.”
Research attracts funding
Spence is the principal investigator and Fromme, Doak, Weierstall and Schmidt are co-investigators on a recent grant from the National Science Foundation to study the dynamics of electron transfer. The major goal of the grant is to unravel the conformation changes involved in the inter-protein electron transfer between the membrane protein complex photosystem I and the soluble electron acceptor protein ferredoxin by light induced time resolved femtosecond crystallography.
Fromme is leading another team that includes Spence, Doak, Weierstall, Schmidt and Alexandra Ros, assistant professor in chemistry as recipient of a new $1.46 million grant from the National Institutes of Health to further study femtosecond nanocrystallography of membrane proteins. “The aim of the grant is to develop the new method further, which may well revolutionize the field of structural biology. That said, the innovations developed for this grant will surely produce exciting and radically new research activities,” said William Petuskey, chair of ASU’s Department of Chemistry and Biochemistry.
SOURCES:
Petra Fromme, PFromme@asu.edu
John Spence, spence@asu.edu
MEDIA CONTACT:
Carol Hughes, carol.hughes@asu.edu
480-965-6375/office, 480-254-3753/cell

