The emergence of form: Study expands horizons for DNA nanotechnology

Hao Yan directs the Biodesign Center for Molecular Design and Biomimetics (BCMDB) and holds the Milton D. Glick Distinguished Professorship in ASU’s School of Molecular Sciences.
In the world of biomolecules, none is more iconic, nor more versatile, than DNA. Nature uses the famous double helix to store the blueprints of all living forms, drawing on a four-letter alphabet of nucleotides.
Researchers in the field of DNA nanotechnology have been inspired by the seemingly inexhaustible variety of living forms nature has fashioned from this genetic raw material. The field seeks to emulate nature’s creative enterprise and even extend the possibilities of DNA architecture beyond what nature has created.
In a new study, Hao Yan and his colleagues Nicholas Stephanopoulos and Petr Sulc, explore a basic building block used in the fabrication of many DNA nanoforms. Known as a Holliday junction, this nexus of two segments of double-stranded DNA has been used to form elaborate, self-assembling crystal lattices at the nanometer scale, (or roughly 1/75,000th the width of a human hair).
The structures take their name from molecular biologist Robin Holliday, who first proposed their existence in 1964. Holliday junctions play an essential role in nature, where they are involved in a process known as homologous recombination, a driving force in generating novel genetic variation in living things.
Since the inception of DNA nanotechnology, the field has made remarkable strides, using DNA components to design tiny structures of intricate beauty as well as nanoscale devices whose applications touch on fields as varied as photonics, computer storage, biosensing and tissue regeneration.
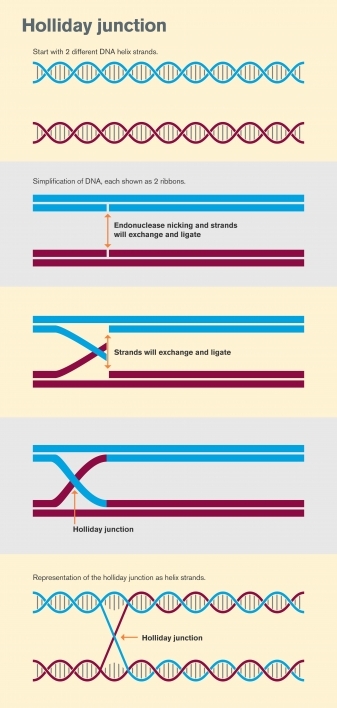
The graphic shows the formation of a Holliday junction from two separate strands of double-stranded DNA. The form is a basic building block used in DNA nanotechnology to form crystal nanostructures. Holliday junctions are also found in nature during cell meiosis and form an intermediate stage during the process of homologous recombination. Graphic by Shireen Dooling
Yan has been at the forefront of the field’s rapid transformations, designing a myriad of useful nanoarchitectural forms, from cartwheeling nanorobots and DNA spiders to cancer-fighting seek-and-destroy devices.
The new study uses crystallography techniques to describe the characteristics of 36 basic variants of the Holliday junction. The results show that the effectiveness of a given Holliday junction for the construction of crystalline nanoarchitectures is sensitively dependent not only on the arrangement of the four nucleotide pairs forming the junction, but also on sequences forming the junction’s four protruding arms. Some DNA sequences act to enhance the crystallization process of these forms, while six of the 36 Holliday junction variants were deemed “fatal” due to their failure to form crystals.
Yan directs the Biodesign Center for Molecular Design and Biomimetics and holds the Milton D. Glick Distinguished Professorship in ASU’s School of Molecular Sciences. Stephanopoulos and Sulc are also faculty members in center and school.
The research findings, which represent the first systematic study of Holliday junctions, recently appeared in the journal Nature Communications.
DNA proves to be an ideal material with which to design and fabricate nanoscale structures. The consistent and predictable nature of base pairing among DNA’s four nucleotides ensures that properly engineered forms will reliably self-assemble into desired structures. To this end, various elaborate nanoforms have been constructed using fundamental DNA building blocks, one of the most popular and useful being the Holliday junction. DNA crystals composed of repeating structural units are key ingredients for nanotechnology applications, allowing for versatile and scalable design features.
Holliday junctions are observed in nature as an intermediary stage during the process of cell meiosis. The result of this transformation is an exchange of genes between maternal and paternal chromosomes. This process, known as homologous recombination, occurs in four stages. (See illustration at right.)
First, a pair of double-stranded DNA helices sit alongside each other. An enzyme known as an endonuclease then causes a single-stranded break in each of the two double strands. The next step, known as strand invasion, occurs when the free ends of each single-stranded break join together, causing the originally separate double strands to be intertwined.
This cross-shaped structure, which joins the two separate double strands of DNA together, is the Holliday junction. In biological processes, the junction is then “resolved” when another enzyme cuts the Holliday junction in one of two ways, both resulting in two separate DNA strands, which differ from the original strands because the Holliday junction has introduced new DNA segments into the two DNA double strands.
This form of DNA recombination is a universal biological event of great importance. It simultaneously acts to preserve genome integrity through mechanisms of DNA repair while generating novel variability, without which, organisms would soon reach an evolutionary dead end. The key structure in the shuffling of the DNA deck during cell division is the Holliday junction.
It was later noted that the Holliday junction motif could be used as a powerful building block for a multiplicity of artificial DNA structures. Although the Holliday junctions occurring during cell division can slide along the DNA length, in a process known as branch migration, the junctions used for constructing DNA nanostructures are immobilized because the sequences flanking them are not complementary.
Chad Simmons
“The first immobile Holliday junction was described in 1982, and this sequence has since been used exclusively in self-assembling DNA crystals,” said Chad Simmons, first author of the paper and the lead scientist applying X-ray crystallography for this study. “Our work sought to change this paradigm by probing the 35 other possible immobile junctions. As a result, we were able to identify several sequences that yielded superior performance compared to their predecessors in terms of their ability to robustly crystallize and diffract to high resolution, and which allowed the ability to control the symmetry of the lattice arrangement.
"This required an exhaustive effort that yielded 134 new crystal structures, and we are very excited to share a comprehensive toolbox of sequence combinations to direct the design and construction of future self-assembling DNA crystal systems.”
The new research demonstrates that most Holliday junction variants produce self-assembling crystals, though six fatal junction arrangements were incompatible with crystal formation. The common feature in these failed junctions was their lack of two critical binding sites for ions, which are essential for crystal formation.
Nicholas Stephanopoulos
“This study was fascinating because it showed how subtle variations in Holliday junction geometries — which could be understood at the single-nucleotide level — could have dramatic effects on crystal assembly and symmetry. This is truly 'molecular science,' allowing us to eventually engineer interactions at the molecular level that will give rise to exciting nanomaterials with unprecedented control,” Stephanopolous said.
“One of the challenges of this research was to determine why some Holliday junctions could produce crystals, but others do not. Empirically, we could study the crystal structures of those junctions that crystallize, but to understand the behavior of the fatal junction arrangements that do not, computational chemistry was needed,” Sulc said.
Petr Sulc
"To this end, we teamed up with Dr. Miroslav Krepl and Professor Jiri Sponer from the Czech Academy of Sciences, who simulated all Holliday junctions at atomistic resolution, and gained the critical insight that the fatal junctions were not able to bind ions that stabilize conformation. This effort provided an excellent example of where computer modeling and experiments can jointly explain complex phenomena,” he said.
The new research provides valuable clues for the design and development of novel forms to be added to the ever-growing plethora of nanostructures and nanodevices serving a broad range of applications in electronics, imaging, computer science and medicine.
More Science and technology

How AI is changing college
Artificial intelligence is the “great equalizer,” in the words of ASU President Michael Crow.It’s compelled industries, including…

The Dreamscape effect
Written by Bret HovellSeventh grader Samuel Granado is a well-spoken and bright student at Villa de Paz Elementary School in…

Research expenditures ranking underscores ASU’s dramatic growth in high-impact science
Arizona State University has surpassed $1 billion in annual research funding for the first time, placing the university among the…