ASU and USF investigators collaborate to explain where DNA repairs occur most frequently

ASU researchers Marcia Levitus, associate professor in the School of Molecular Sciences and the Biodesign Institute, and Wade Van Horn, assistant professor in the School of Molecular Sciences and investigator with the Biodesign Institute's Center for Personalized Diagnostics and the Magnetic Resonance Research Center.
From hair and eye color to how our biological system is regulated, the blueprint of life is held in the genome.
The gene also is what provides the instructions to the DNA as to what proteins to make and when, but our DNA is under constant attack from our environment — sun exposure, the air we breathe, the foods we eat and the body’s own metabolic processes to sustain life. If the enzymes that repair DNA are not signaled, damaged DNA may affect cell division and may result in the formation of tumors that can lead to cancer.
For a genome to survive, repairs to the DNA base pair is crucial. But at what rate and where do these repairs occur most frequently?
Marcia Levitus, associate professor in the School of Molecular Sciences and the Biodesign Institute at Arizona State University, received a National Science Foundation grant to lead a study on the dynamics of DNA sequence and deformability on lesion recognition and excision in the base excision repair pathway, which will help us understand key aspects of DNA repair, DNA mutation rates and molecular evolution.
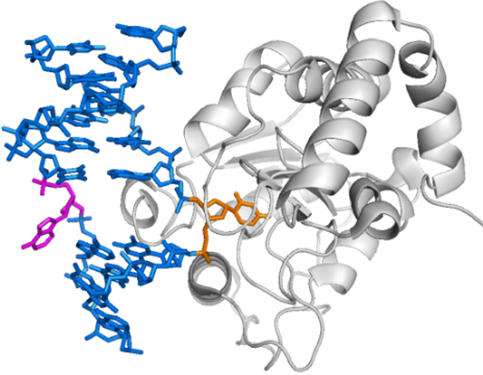
“Thousands of spontaneous lesions occur to a cell’s DNA on a daily basis, and to maintain the integrity of their genomes, cells have evolved mechanisms to repair damaged DNA," said Levitus. "The mechanism responsible for removing small lesions from the genome is the base excision repair pathway. This pathway is responsible for removing most oxidized, alkylated and deaminated nucleobases from the genome, and is initiated by specialized DNA glycosylases that catalyze the excision of the damaged base.”
Each of us has approximately 3 billion base pairs of DNA, and the instructions for building, repairing or maintaining itself is determined from the base sequence. The average person’s cells are estimated to have 70,000 DNA lesions. It would seem impossible for a cell to maintain the integrity of the DNA or survive if it didn’t evolve or adapt to the surrounding conditions it is continually subjected to.
During a cell cycle, a series of steps or events lead up to DNA replication known as a cell checkpoint, where cells are evaluated for ideal criteria before they can begin the replication process. The mechanisms for a biological system have a critical role in order to assure the genetic material being passed on is correct. Our DNA is constantly undergoing replication and division, and as in any system, errors can happen in the process. The majority of errors in our DNA are able to be detected in the cell cycle through very efficient repair systems, and the human body remains healthy.
The code of instructions in the DNA consists of four chemical bases: adenine (A), guanine (G), cytosine (C) and thymine (T). Nucleic acids are formed from nucleotides. In DNA, there are three parts to a nucleotide: a deoxyribose (a five carbon sugar molecule), a phosphate group and a nitrogenous base. DNA base pairs to form units are made up in the sequence A with T and C with G.
Role of DNA sequence and deformability on lesion recognition and excision in the base excision repair pathway
Levitus stated recent studies have shown that a number of DNA repair enzymes are more active on particular sequences of DNA than others.
“This sequence specificity results in nonrandom patterns of repair that can lead to nucleotide positions with lower repair efficiencies and exceptionally high mutation frequency,” said Levitus. “Identifying and characterizing factors that determine the rates and spectra of spontaneous mutations is a critical step toward understanding molecular aspects of the evolutionary process.”
Collaborating with Levitus on this research are Wade Van Horn, co-principal investigator and associate professor at ASU's School of Molecular Sciences, investigator at the Biodesign Institute, the Center for Personalized Diagnostics and the Magnetic Resonance Research Center, and Arjan van der Vaart, an associate professor and associate chair at the University of South Florida's Department of Chemistry.
“DNA carries the instructions that govern biology, and there are a variety of ways in which these DNA instructions can be changed (mutated). Consequently, biology has several diverse ways to identify and fix these DNA alterations,” said Van Horn. “Our collaborative studies seek to understand how biology recognizes these changes (lesions), particularly by investigating the relationship between the DNA sequence, its dynamics (i.e., movement) and the efficacy in identifying DNA lesions. Our team will use a mix of cutting edge computational and experimental approaches to investigate how biology protects the genetic instructions carried by DNA.”
The main goal of this project is to elucidate the molecular basis for sequence effects in lesion repair by DNA glycosylases. Levitus, Van Horn and van der Vaart will test the hypothesis that damage that occurs within a rigid region of DNA is repaired more slowly than damage that occurs within a flexible region. Elucidating the molecular basis for sequence effects in DNA repair requires a multipronged approach that takes advantage of the strengths of a variety of experimental and computational approaches. This collaborative research will capitalize on the expertise in DNA biophysics (Levitus and van der Vaart), fluorescence spectroscopy (Levitus), NMR spectroscopy (Van Horn) and molecular dynamics (MD) simulations (van der Vaart).
More Science and technology

Hidden viruses thrive in desert wildlife
As the sun rises over the Sonoran Desert, bright green lovebirds gather noisily around backyard feeders. At dusk in the Arizona…

2 ASU faculty named NAI fellows
Professors Krishnendu Chakrabarty and Rosa Krajmalnik-Brown are among the latest Arizona State University faculty members to be…

How AI is changing college
Artificial intelligence is the “great equalizer,” in the words of ASU President Michael Crow.It’s compelled industries, including…