X-ray laser study identifies crystalline intermediate in our 'pathway to breathing'

(From left) Austin Echelmeier, Alexandra Ros, Petra Fromme and Raimund Fromme, all from ASU’s School of Molecular Sciences and the Biodesign Institute’s Center for Applied Structural Discovery.
Scientists from Arizona State University’s School of Molecular Sciences, in collaboration with colleagues from Albert Einstein College of Medicine in New York City, have captured for the first time snapshots of crystal structures of intermediates in the biochemical pathway that enables us to breathe.
Their results, published today in the Proceedings of the National Academy of Sciences in the article "Snapshot of an Oxygen Intermediate in the Catalytic Reaction of Cytochrome c Oxidase," provide key insights into the final step of aerobic respiration.
“It takes a team to conduct such a sophisticated experiment,” said Associate Professor Alexandra Ros who, together with her graduate student Austin Echelmeier and former intern Gerrit Brehm, developed the hydrodynamic focusing mixer that made these experiments possible.
The mixer is a microfluidic device, which is high-resolution, 3D-printed and enables two streams of oxygen-saturated buffer to mix perfectly with a central stream containing bovine cytochrome c oxidase (bCcO) microcrystals. This initiates a catalytic reaction between the oxygen and the microcrystals.
In the beginning
This research was instigated by a conversation between Professor Petra Fromme, director of the Biodesign Institute’s Center for Applied Structural Discovery (CASD); Raimund Fromme, School of Molecular Sciences associate research professor; and Professor Denis Rousseau from the Albert Einstein College of Medicine in New York City who works on the structure of cytochrome c oxidase, a key enzyme involved with aerobic respiration.
Cytochrome c oxidase (CcO) is the last enzyme in the respiratory electron transport chain of cells located in the mitochondrial membrane. It receives an electron from each of four cytochrome c molecules, and transfers them to one oxygen molecule (two atoms), converting the molecular oxygen to two molecules of water.
Researchers at CASD, including ASU’s Richard Snell Professor of Physics John Spence, helped to pioneer a new technique called time-resolved serial femtosecondA femtosecond is a millionth of a billionth of a second. crystallography (TR-SFX). This technique takes advantage of an X-ray Free Electron Laser (XFEL) at the Department of Energy's SLAC National Accelerator Laboratory at Stanford University.
TR-SFX is a promising technique for protein structure determination, where a liquid stream containing protein crystals is intersected with a high-intensity XFEL beam that is a billion times brighter than traditional synchrotron X-ray sources.
While the crystals diffract and immediately are destroyed by the intense XFEL beam, the resulting diffraction patterns can be recorded with state-of-the-art detectors. Powerful new data analysis methods have been developed, allowing a team to analyze these diffraction patterns and obtain electron density maps and detailed structural information of proteins.
The method is specifically appealing for hard-to-crystallize proteins, such as membrane proteins, as it yields high-resolution structural information from small micro- or nanocrystals, thus reducing the contribution of crystal defects and avoiding tedious (if not impossible) growth of large crystals as is required in traditional synchrotron-based crystallography.
This new “diffraction before destruction” method has opened up new avenues for structural determination of fragile biomolecules under physiologically relevant conditions (at room temperature and in the absence of cryoprotectants) and without radiation damage.
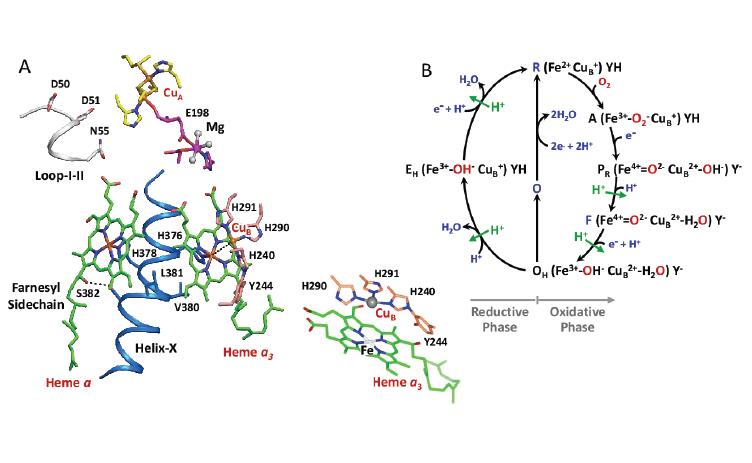
CcO reduces oxygen to water and harnesses the chemical energy to drive proton (positively charged hydrogen atom) relocation across the inner mitochondrial membrane by a previously unresolved mechanism.
In summary, the TR-SFX studies have allowed the structural determination of a key oxygen intermediate of bCcO. The results of the team’s experiments provide new insights into the mechanism of proton relocation in the cow enzyme as compared to that in bacterial CcOs, and paves the way for the determination of the structures of other CcO intermediates, as well as transient species formed in other enzyme reactions.
Other coauthors on this paper, not previously mentioned, include Izumi Ishigami, Ariel Lewis-Ballester and Syun-Ru Yeh, all from the Albert Einstein College of Medicine; Nadia Zatsepin and Stella Lisova of the ASU Department of Physics; Jesse Coe, Zachary Dobson, Garrett Nelson and Shangji Zhang all from the School of Molecular Sciences and CASD; Thomas Grant from University at Buffalo, State University of New York; and Sébastien Boutet, Raymond Sierra and Alexander Batyuk all from SLAC.
This research was also supported by an NIH R01 (Petra Fromme) and the NSF BioXFEL STC.
More Science and technology

How AI is changing college
Artificial intelligence is the “great equalizer,” in the words of ASU President Michael Crow.It’s compelled industries, including…

The Dreamscape effect
Written by Bret HovellSeventh grader Samuel Granado is a well-spoken and bright student at Villa de Paz Elementary School in…

Research expenditures ranking underscores ASU’s dramatic growth in high-impact science
Arizona State University has surpassed $1 billion in annual research funding for the first time, placing the university among the…